 Please note that using the default 'All' ASL expression, the program will align SSEs and at the same time minimize RMS deviation of the C-alphas, for which you will have to apply a residue-based ASL for the alignment (rather than atom based).įor example, here the ASL 'a.pt CA' (for C-alpha alignment) fails, whereas the ASL 'backbone' would work for the same system. In Tools → Superposition, you can use the ASL expression 'backbone' to superimpose based on the backbone, or 'a.pt CA' to superimpose based on C-alpha only. $SCHRODINGER/utilities/structalign -force The TM-align will first find the best equivalent residues of two proteins based on the structure similarity and then output a TM-score. Ii) Align the proteins from the command-line, using the following command: 300-400 Furthermore, you may wish to restrict the alignment to just the backbone atoms, so you can say: align structure2 and resi 1-100 and name n+ca+c+o, structure1 and resi 300-400 and name n+ca+c+o or in short form: align structure2 & i. I) Use a subset of the two proteins for the alignment, using the 'Select' option in the alignment panel. In the command-line window (Depending on your PyMOL version, Windows labels this Tcl/Tk GUI or The PyMOL Molecular Graphics System), type the following commands: select AB, chain A+B hide all show cartoon, AB orient AB Note that in the panel at the top right, you now can operate on the subset AB using the buttons. Structures were aligned in PyMol using backbone heavy atoms (CA, C. The alignment code looks at secondary structure elements (SSE) for aligning two proteins, so there has to be at least some similarity in SSEs in order for the structures to be aligned.ģ) In rare cases, it has been found that the alignment code does not align systems with multiple chains. 2.1 RMSD values capturing variations in peptide backbone and side chain confor. 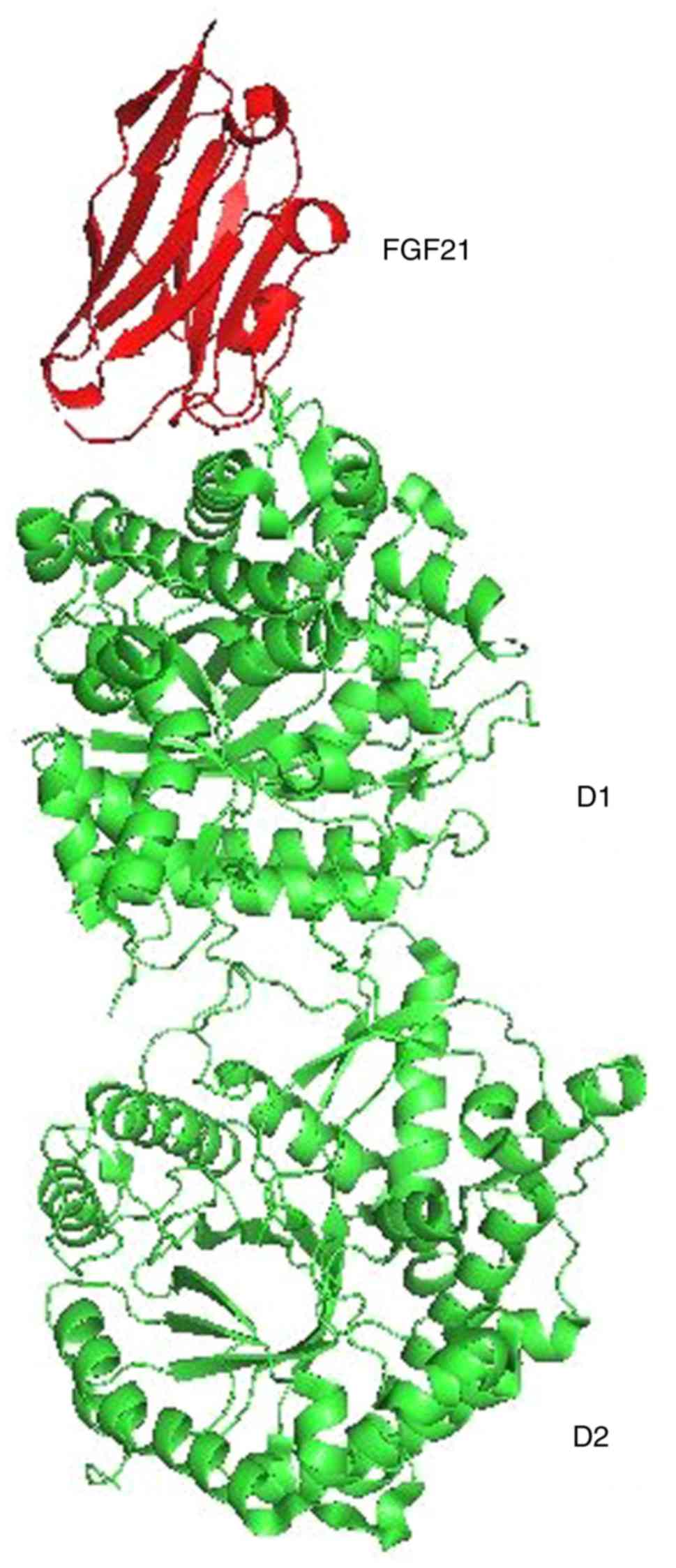
The backbone and side-chain resonance assignment experiments were. First, we will add hydrogens into the structure in PyMOL and then show the backbone H-bond distances. To visualize these \bonds', we can measure the lengths between groups of atoms, speci cally where polar contacts occur. 
1) Ensure that all the structures being aligned are included in the Workspace (In box checked) and not just highlighted (selected) in the Project Table.Ģ) Check that the structures being aligned are similar. 0.5mM HRGsingle peptide was titrated into 0.05mM of sHIPwt, or sHIPqp respectively. peptide backbone, which tends to be attracted to the partially negative oxygens in the backbone. However, each residue is distinguished by a side chain of atoms that is not on the backbone, but is attached to the alpha carbon backbone atom, denoted C, for. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
